Modern Biological AI
Rapid advances in this space rivalled only by NLP advances. This is about what deep learning adds.
Protein Language Models
They’re in a sequence, why not toss an NLP on them? See other notes: this is not language! Asymmetry here.
ESM (Evolutionary Scale Modeling) models from Facebook are a family of protein language models. Train a transformer on protein sequences using BERT’s “masked prediction objective” (TODO) — but do this on the 20 AA alphabet. Same idea as BERT → ClinicalBERT → ESM-2 but on proteins. Learns (claim) evolutionary constraints, coevolutionary patterns, and functional motifs. Imagine a string and coil it. This is asking “are AA’s in contact” based on a cross-axis of sequences (one of the graphs). This is called “Contact Recovery”.
Rao et al — which protein sequences are “attending” to each other (using Transformers).
Variant Effect Prediction: If I change something in DNA, is the change in the protein pathogenic or not?
What Protein LMs get wrong:
- Note that Plausibility Pathogenicity!
- Conservation Function! If a position is conserved evolutionarily, it doesn’t mean that it is important biologically. E.g. Selfish sequences.
- You can have underrepresented proteins: viral, synthetic, orphan.
- Context is important: different tissue, different species, different pathway.
- MSA (Multiple Sequence Alignment) methods outperform: LMs are based on sequence and MSAs bring evolution to bear.
ESM assumes sequential generative process. Makes sense for language but not for protein sequences — driven by evolution which is a branching process. So ESM captures marginal distro but not the joint across homologous sequences.
Here’s a combo: An MSA Transformer! So row attention can capture sequence context and column attention can capture “coevolutionary coupling” (Rao et al, 2021). You then get a coevolution signal in addition to “per-residue embeddings”.MSA Transformer showed that you can recover contacts better than single-sequence LMs (ESM) or classical DCA.
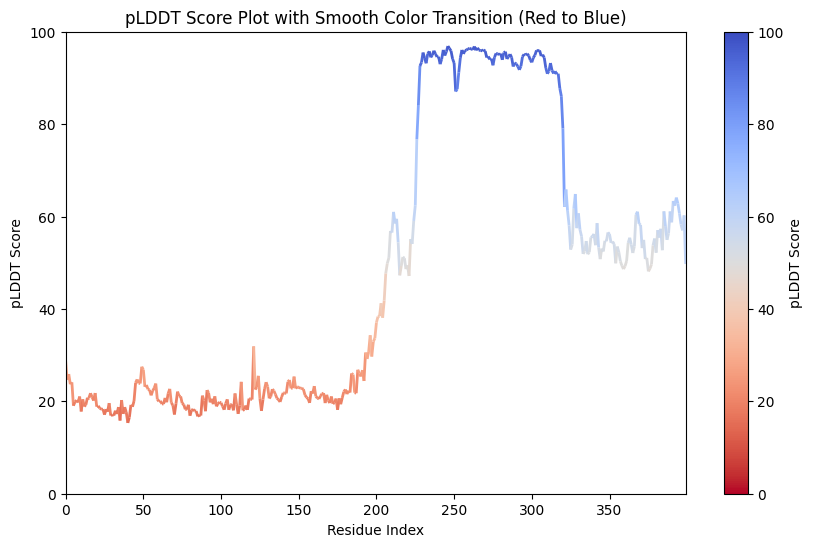
Now AlphaFold 2’s “Evoformer” has the same axial attention, then adds a pair representations (residue x residue matrix), adds structural supervision (trained end-to-end to predict 3D coordinates and not just contacts.) It gets a lot of things right and wrong. Nothing’s perfect. See pLDDT scores (pLDDT accuracy, residue-level metric, local; there’s also TM Score which is more global, might be better for overall 3D structure.)
Genomic Sequence Models
Goal here is to predict gene expression and regulatory effects directly from DNA sequence without GWAS/eQTL (latter is about mRNA sequences; “variant effect prediction” TODO)
THis is a hard problem because of ~3.2B base pairs and regulatory elements can be 1 megabase from genes! If you toss this at a standard transfomer, you run into ye olde problems and will need reaaaaaaly long contexts.
Important: Regulatory function depends on a lot more than mere sequence — cellular context like tissue, chromatin state, development state, etc.
If there’s a key takeaway it is this. ‘Raw’ Sequence alone ain’t enough. DNA and protein sequences don’t exist in a vacum. Context is everything.
Lots of amazing work here but: do they learn biology? To date, not too many exciting results. State of things (individual opinion):
| Task | Perf | Clinical Readiness |
|---|---|---|
| Expression from Reference | Strong | Low |
| Variant effect (common) | moderate | Low |
| Variant effect (rare) | Weak | Not ready |
| Regulatory discovery | Promising | Research Only |
3D Molecular Modeling
Why? The same 2D graph can have different 3D conformations. You get things like interatomic distances, all sorts of angles (bonds, dihedral).
So some constraints here:
- Scalar stuff/predictions (energy, binding affinity) are invariant
- Vector predictions (forces, dipole movement) are equivariant
Take a molecule, any molecule. Now number its atoms. Then make an Adjacency Matrix (1 if there’s a bond between two indices; 2 if there’s a double bond.) This will just capture bond features. The actual atoms would be in a different vector. Now with the Adjacency Matrix, you can use path lengths and all that but this doesn’t capture distances. Now you can have a matrix of distances and even one of angles (some will even use coordinates) but even that is not enough to determine structure. An open problem in research. See also the symmetries handled by each approach.
The Inverse Problem
Reverse what you’re doing (“I want to predict structure or properties from sequence”) to “I want the protein to do X, predict the sequence (or structure).” — You’re designing proteins. Now think about this in terms of causal inference. Some rough mapping there (“what input would produce this outcome?”)
Say you want a brighter than GFP.
E.g. ProteinMPNN is a DL model for design. 52% per residue recovery. Give it a 3D backbone and it will give you a sequence.
People are using diffusion methods too like RFDiffusion.
In the end the output of a design model is a hypothesis, not a protein!
Random
Think about the inverse modeling. If you give the model a number (e.g. GFP brightness) and say “What sequence generated this?” you’ll get a confused model (“There’s like a lot of sequences bro.”). But that uncertainty is very helpful to the person asking the question!
Geometry and Evolution are increasingly important.